Computational Genetics and Evolution @UNM
Our research develops theory, computational approaches, and statistical tools to uncover mechanisms of rapid evolution in changing environments. As conditions in nature change rapidly, such as seasonal changes, adaptation is far more likely to occur if newly adaptive alleles are already present in the population in the form of standing genetic variation. Therefore, our focus is on understanding mechanisms that promote genetic variation in changing environments and how they shape patterns of rapid evolution. In our lab, we tackle the issue from several perspectives: (1) rapid molecular evolution at one or few genetic loci, (2) rapid phenotypic evolution in polygenic traits, and (3) rapid evolution of whole communities. In addition, we develop approaches to infer the genetic basis of complex trait variation in humans.
(1) Our theoretical work is currently focusing on evolutionary mechanisms of rapid molecular adaptation in temporally changing habitats, the mechanisms of rapid molecular adaptation during niche expansions, and the evolution of plasticity and canonization in structured populations.
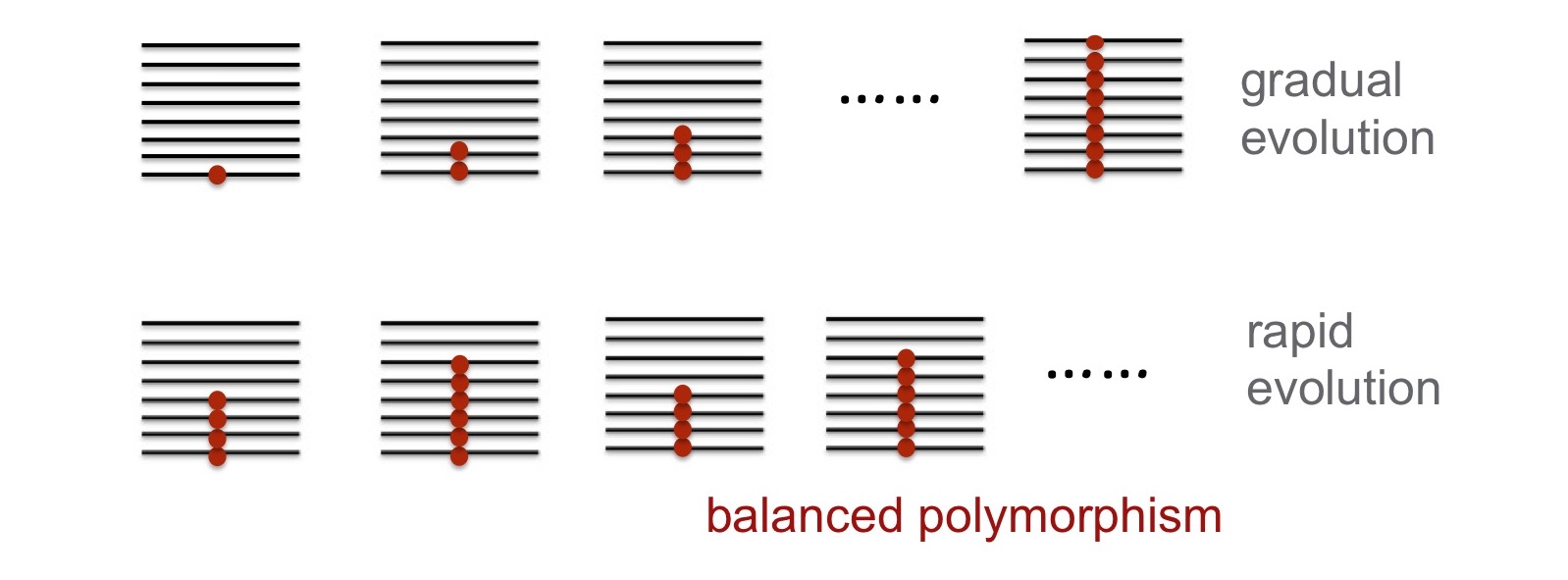
(2) We are developing theory and statistical approaches for inferring genetic basis of complex phenotypes using whole-genome markers in humans. We are also using computer simulations to uncover evolutionary mechanisms of rapid polygenic adaptation.
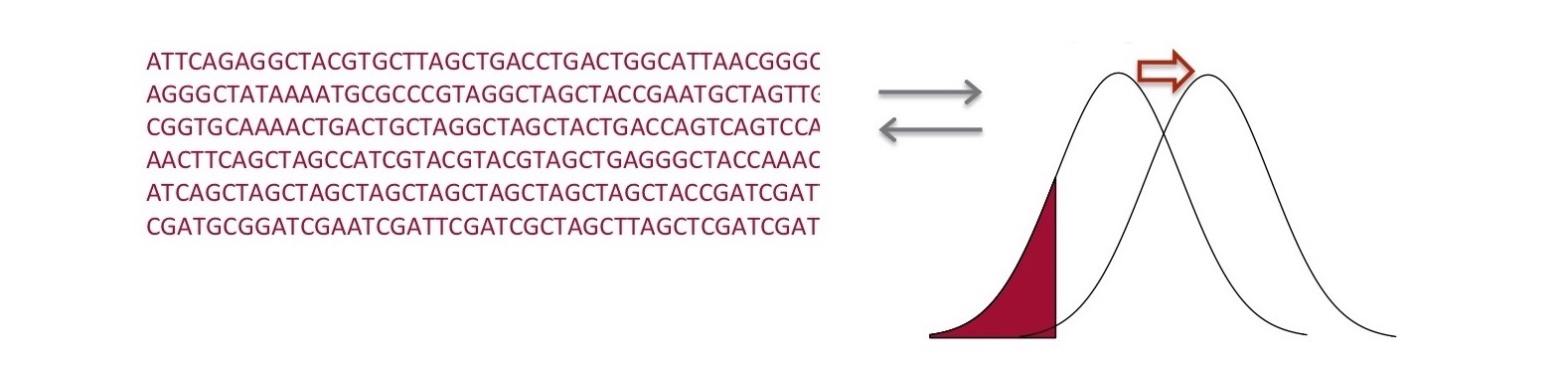
(3) Microbiomes are quintessential complex systems, whose structures are shaped by networks of ecological and evolutionary processes across levels of the organization, from genes within individuals within microbial communities to dynamic host-populations/habitats. We are developing a general framework for eco-evolutonary dynamics of microbiomes that will allow us to simultaneously evaluate a myriad of interactive processes that shape the patterns of microbiome diversity.
